What are protein interaction networks?
Protein-protein interactions are the representations of physical contacts between proteins in the cell which are specific, occur between defined regions of protein, and have a specific biological function. The knowledge from protein interaction network can be used to characterize the relationship between proteins forming complexes and learn more about the details of signaling pathways [1]. STRING is a database of known and predicted protein-protein interactions which includes direct (physical) and indirect (function) interactions [2].
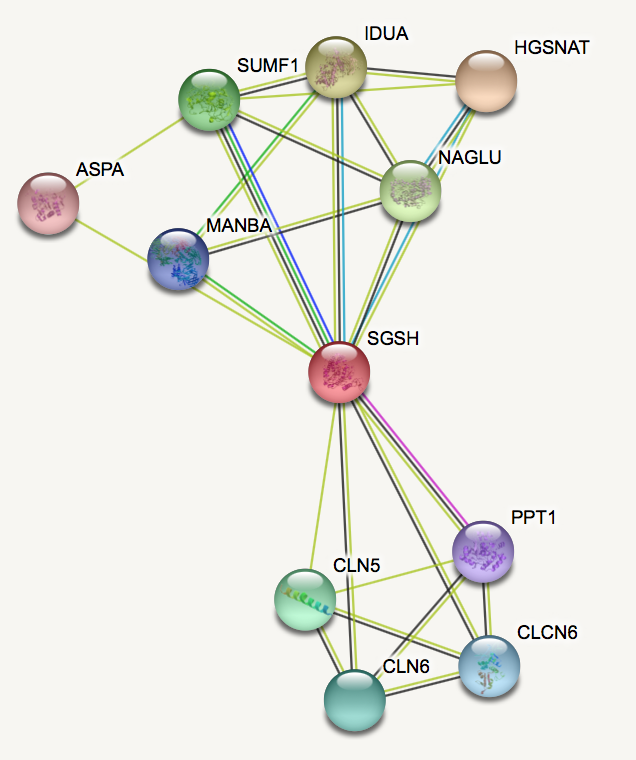
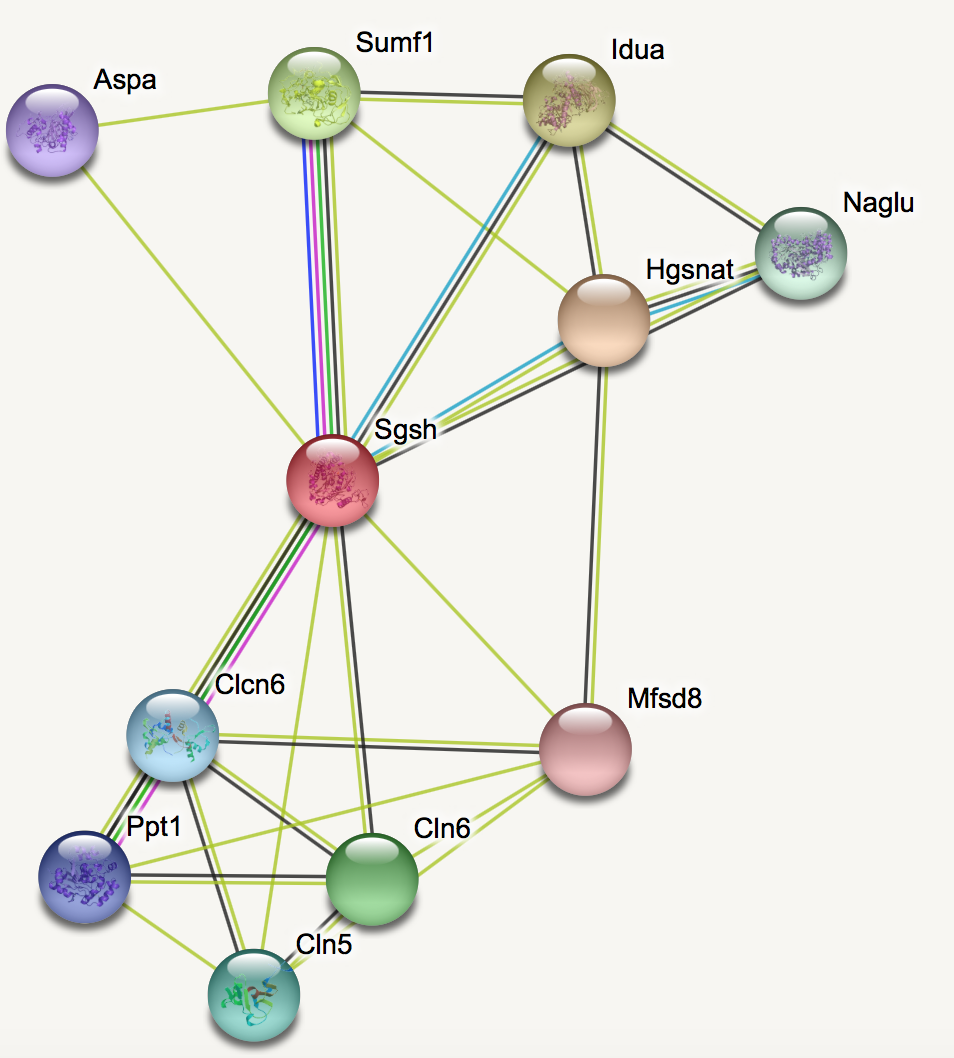
Interaction networks for SGSH was built for humans and mice using the STRING database.
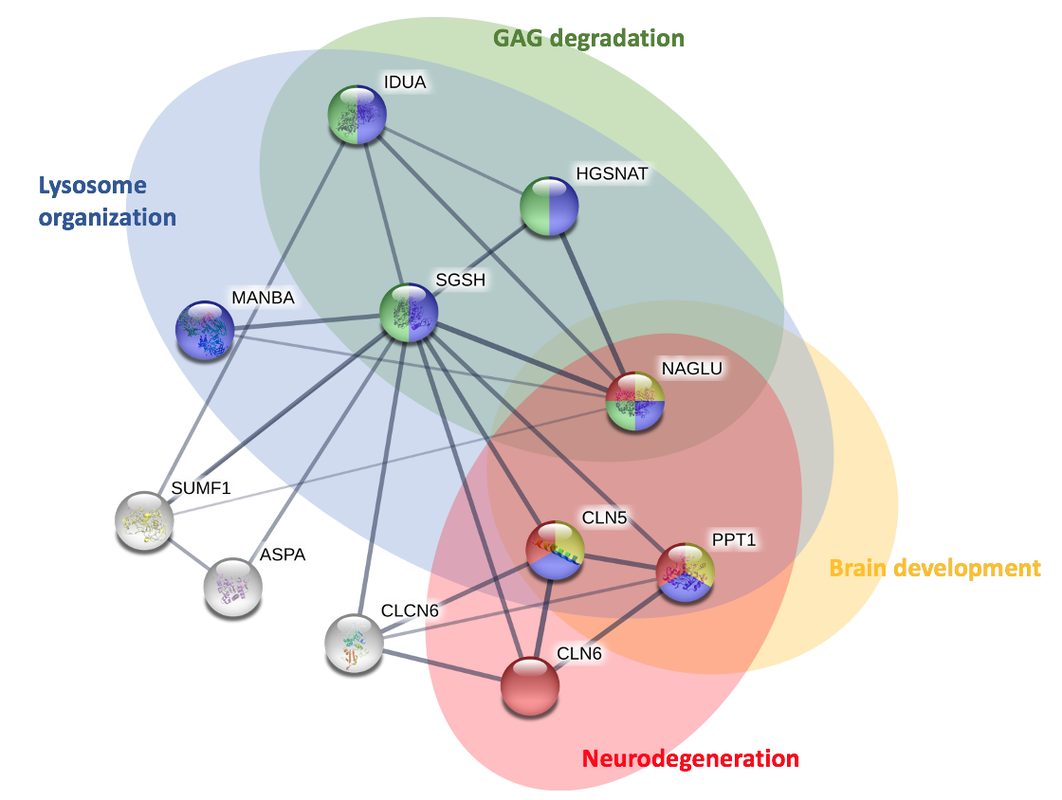
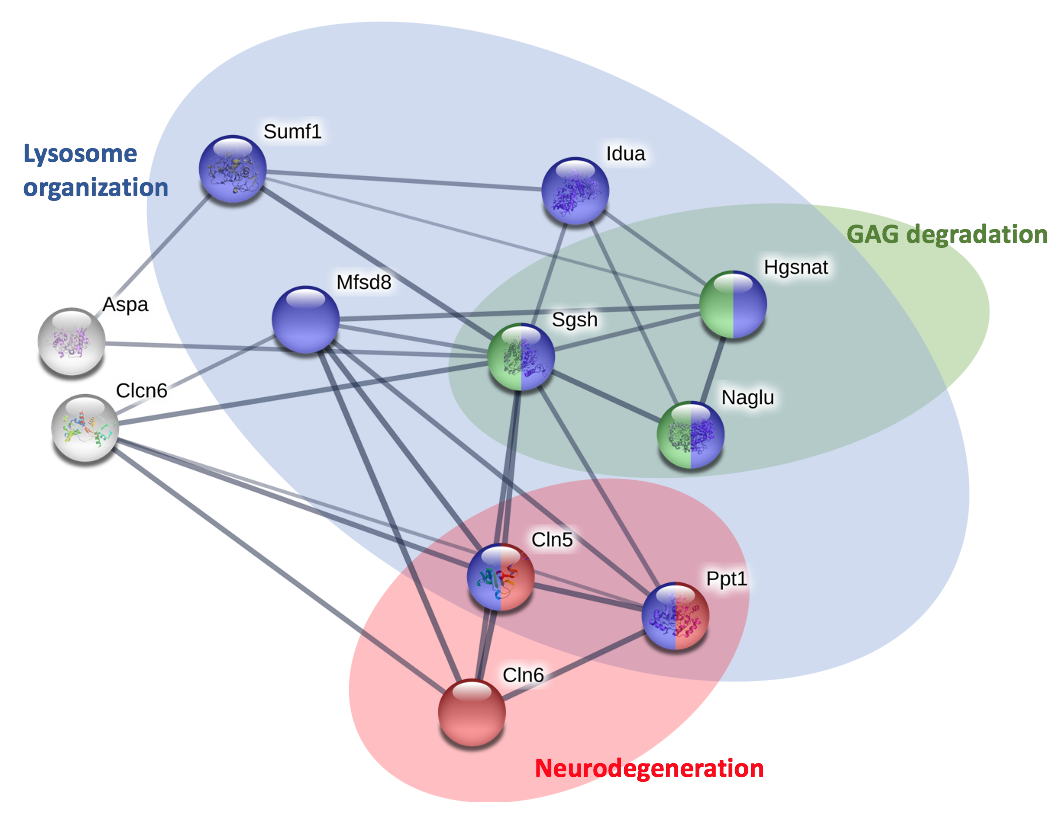
To further analyze the interaction network, gene ontology was then utilized to identify and categorize the main ontologies associated with the proteins.
Discussion
Looking at the protein interactions, it is clear that there are some proteins involved in brain development and neurodegeneration that interact with SGSH in humans. Similar interaction network is seen in mice, suggesting there is possible function in brain development.
References
Header: https://www.interaction-design.org/literature/article/how-to-display-complex-network-data-with-information-visualization
[1] https://www.ebi.ac.uk/training/online/course/network-analysis-protein-interaction-data-introduction/protein-protein-interaction-networks
[2] https://string-db.org/cgi/about.pl
[1] https://www.ebi.ac.uk/training/online/course/network-analysis-protein-interaction-data-introduction/protein-protein-interaction-networks
[2] https://string-db.org/cgi/about.pl
This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.